Virus Surge
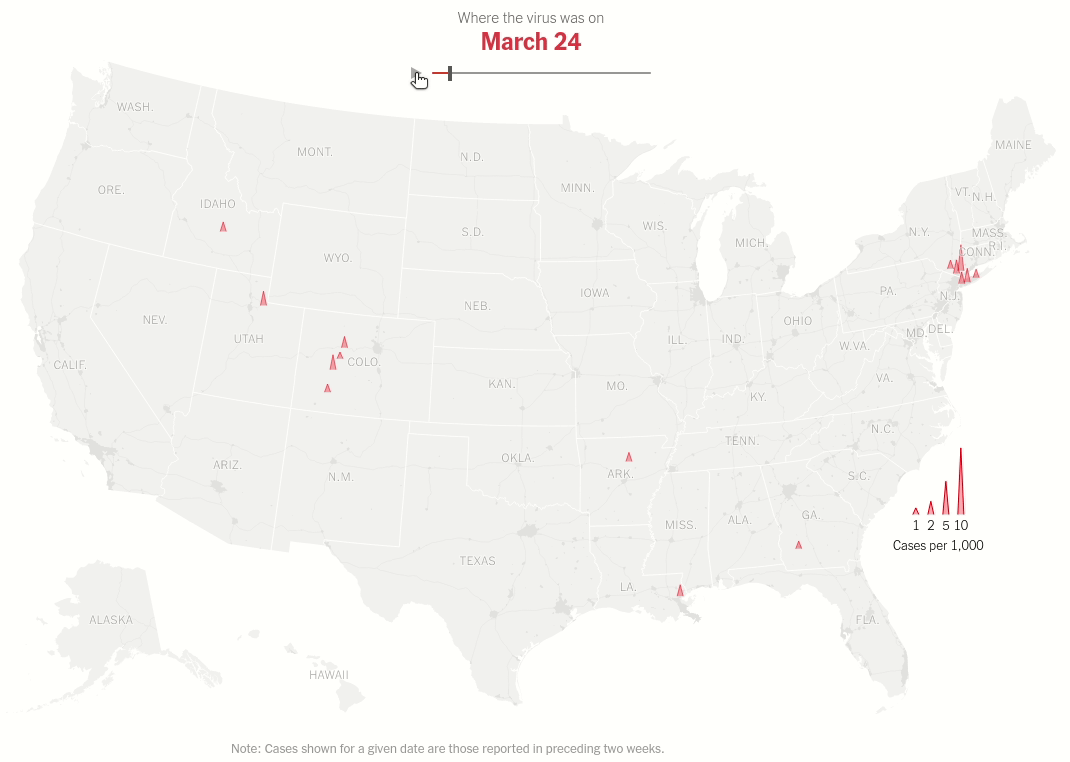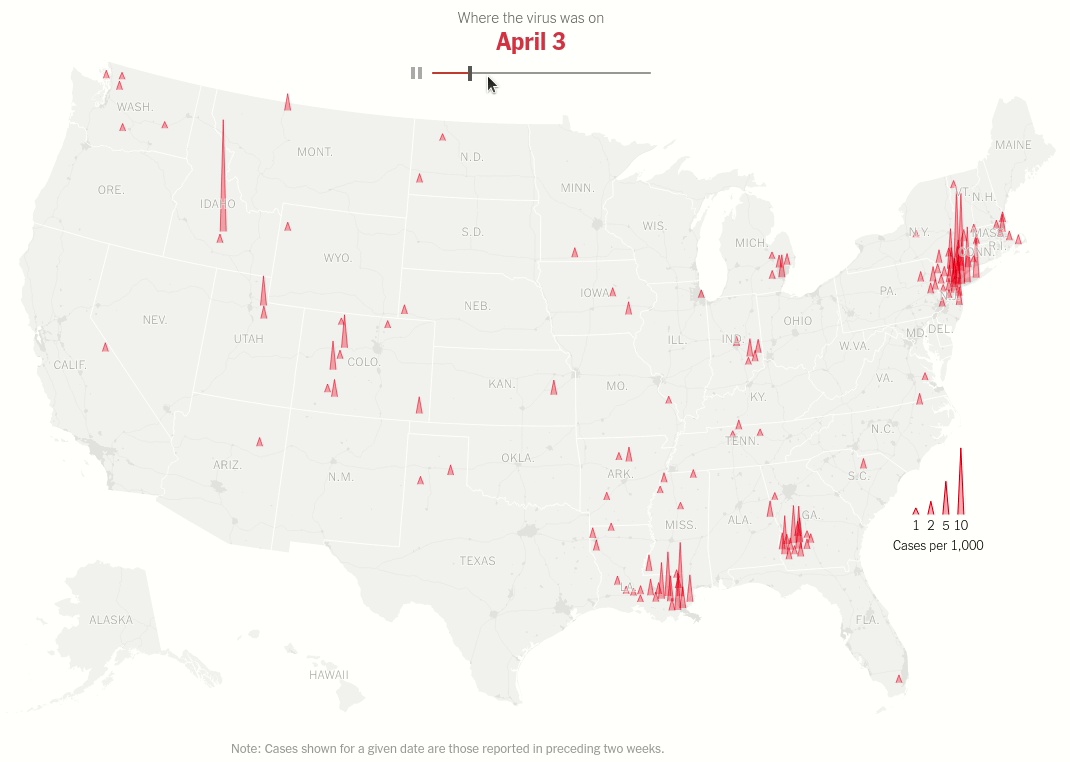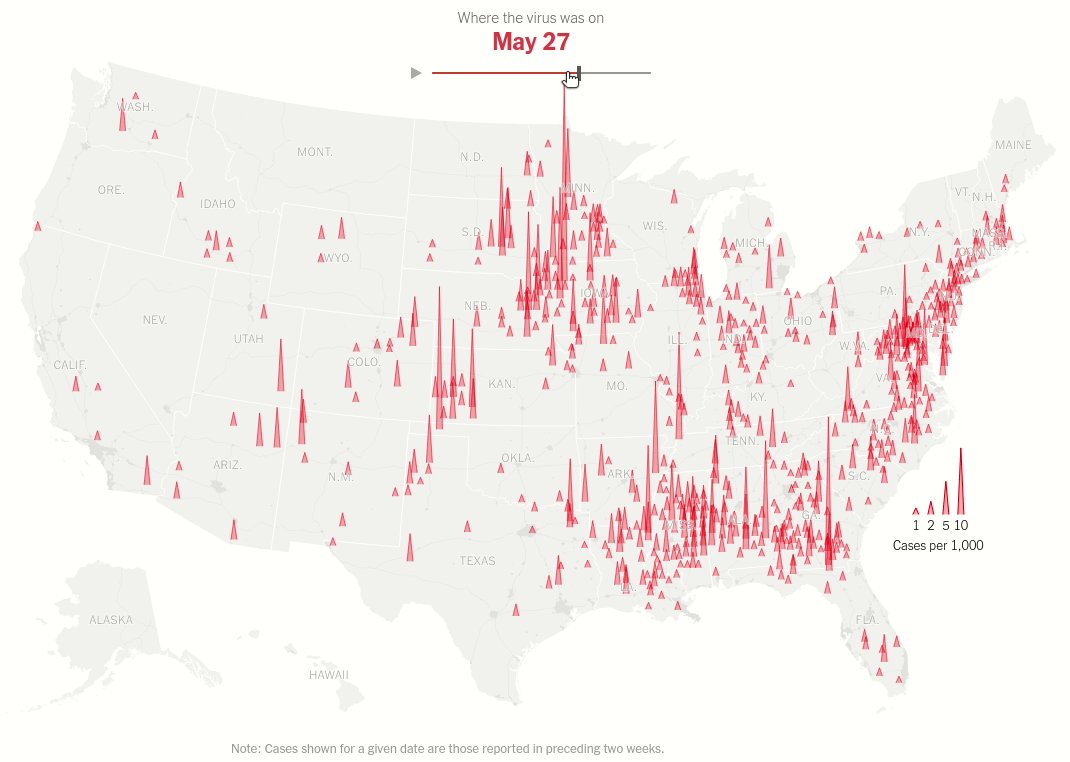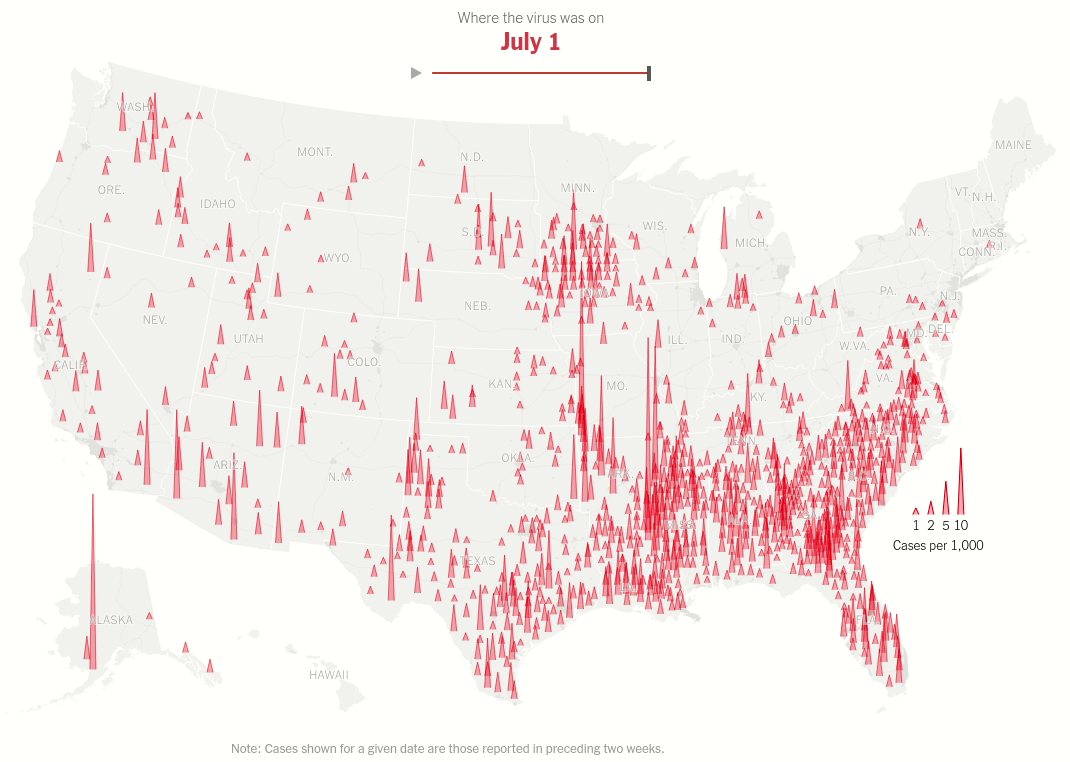
Interactive map with range selector (slider) showing the spread of virus hotspots across the USA over time.
Today we will make this brilliant visualization in the NYT Article U.S. Virus Cases Climb Toward a Third Peak. Both the static visualization and the interactive range slider.
import pandas as pd
import geopandas as gpd
import altair as alt
alt.renderers.set_embed_options(actions=False)
Shapefiles for US States from the topojson/us-atlas repository. Shapefiles from US Census are fine too.
states_shp_uri = 'https://cdn.jsdelivr.net/npm/us-atlas@3/states-10m.json'
states = gpd.read_file(states_shp_uri)
states['lon'] = states['geometry'].centroid.x
states['lat'] = states['geometry'].centroid.y
We will remove the states that are not supported in the albersUsa projection.
# Albers USA projection does not support the following.
states = states[~((states['id'] == '69') | (states['id'] == '78') | (states['id'] == '60') | (states['id'] == '72') | (states['id'] == '66'))]
states.head()
Initial Visualization
alt.Chart(states).mark_geoshape().encode().project('albersUsa')
Getting the COVID data as usual from NYT GitHub Repository
us_covid_data_uri = 'https://github.com/nytimes/covid-19-data/blob/master/us-counties.csv?raw=true'
raw_data = pd.read_csv(us_covid_data_uri)
raw_data['date'] = pd.to_datetime(raw_data['date'])
covid = raw_data.copy()
covid.head()
We will also load the shapefile for counties because we need them for calculating the longitude and latitudes of the centroids of polygons of the counties
counties_shp_uri = 'https://cdn.jsdelivr.net/npm/us-atlas@3/counties-10m.json'
counties = gpd.read_file(counties_shp_uri)
counties.head()
Removing any empty geometries beacause that will give errors when calculating longitudes and latitudes from the centroid
counties.geometry.is_empty.any()
empty = counties.geometry.is_empty
counties = counties[~empty]
counties['lon'] = counties['geometry'].centroid.x
counties['lat'] = counties['geometry'].centroid.y
counties.head()
counties.rename(columns={'id': 'fips'}, inplace=True)
counties.fips = counties.fips.astype(int)
The usual NYC aggregation because of the way data is reported by NYT.
# Extract the boroughs in shapefile geodataframe for NYC
boroughs_nyc = counties[counties['fips'].isin(['36005', '36047', '36061', '36081', '36085'])]
boroughs_nyc['State'] = "NYC"
#combined_nyc = boroughs_nyc.dissolve(by='STATEFP')
agg_nyc_data = boroughs_nyc.dissolve(by='State').reset_index()
agg_nyc_data['fips'] = 1
agg_nyc_data['lon'] = agg_nyc_data['geometry'].centroid.x
agg_nyc_data['lat'] = agg_nyc_data['geometry'].centroid.y
agg_nyc_data = agg_nyc_data.drop(columns=['State'])
counties = gpd.GeoDataFrame(pd.concat([counties, agg_nyc_data], ignore_index=True))
counties.tail()
covid = raw_data.copy()
covid = covid[~((covid['state'] == "Puerto Rico")|(covid['state'] == "Virgin Islands")|(covid['state'] == "Guam")|(covid['state'] == "Northern Mariana Islands"))]
# Extract the boroughs in shapefile geodataframe for NYC
boroughs_nyc = counties[counties['fips'].isin(['36005', '36047', '36061', '36081', '36085'])]
boroughs_nyc['State'] = "NYC"
#combined_nyc = boroughs_nyc.dissolve(by='STATEFP')
agg_nyc_data = boroughs_nyc.dissolve(by='State').reset_index()
agg_nyc_data['fips'] = 1
agg_nyc_data['lon'] = agg_nyc_data['geometry'].centroid.x
agg_nyc_data['lat'] = agg_nyc_data['geometry'].centroid.y
agg_nyc_data = agg_nyc_data.drop(columns=['State'])
counties = gpd.GeoDataFrame(pd.concat([counties, agg_nyc_data], ignore_index=True))
# Make fips in covid data for "New York City" as 1 to reflect what we have done above
covid.loc[covid['county'] == 'New York City','fips'] = 1
#covid = covid.merge(counties[['fips', 'lon', 'lat']], on='fips', how='inner')
#covid = covid.drop(columns=['STATEFP', 'geometry'])
# Getting per day statistics on CASES
covid = covid.assign(daily=covid.groupby('fips')['cases'].diff())
#covid = covid[covid['date'] > '2020-05-01'] # Working with data from May
# Getting past 2 weeks data for cases per given date - sum of cases in past 2 weeks
covid = covid.assign(past_2_weeks = covid.groupby('fips')['daily'].apply(lambda case: case.rolling(window=14).sum().shift(1)))
# Keeping only the latest date data
covid = covid[covid['date'] == '2020-10-27']
# Merging population estimate data - we could have done it earlier but what the heck!
## Getting census data
census = pd.read_csv('co-est2019-alldata.csv')
census = census[['STATE', 'COUNTY', 'STNAME', 'CTYNAME','POPESTIMATE2019']] # Keeping only 2019 population estimates
## Constructing FIPS from STATE and COUNTY codes in census itself
census['STATE'] = census.STATE.astype(str)
census['COUNTY'] = census.COUNTY.astype(str)
census['STATE'] = census.STATE.str.pad(2, fillchar='0')
census['COUNTY'] = census.COUNTY.str.pad(3, fillchar='0')
census = census.assign(fips = census.STATE + census.COUNTY)
census.fips = census.fips.astype(int)
census = census.drop(columns=['STATE','COUNTY','STNAME','CTYNAME'])
## Merging the census data with covid data on FIPS
covid = covid.merge(census, on='fips', how='inner')
## Getting the per capita data
covid = covid.assign(past_2_weeks_per_capita = (covid.past_2_weeks/covid.POPESTIMATE2019)*1000)
covid = covid[covid['past_2_weeks_per_capita'].notna()]
# Per Capita Deaths
covid = covid.assign(deaths_per_capita = covid.deaths/covid.POPESTIMATE2019*1000)
# Per Capita Deaths
covid = covid.assign(cases_per_capita = covid.cases/covid.POPESTIMATE2019*1000)
# Finally merging with geometry
plot_data = counties.merge(covid, on='fips', how='inner')
#plot_data = covid
#plot_data = plot_data.drop(columns=['STATEFP', 'daily', 'past_2_weeks', 'past_2_weeks_per_capita', 'deaths_per_capita'])
print(plot_data.info())
plot_data.head()
Making a custom height column with SVG path strings so that we scale the data using that column
plot_data = plot_data.assign(height = plot_data['past_2_weeks_per_capita'].apply(lambda x: f"M -1 0 L 0 -{x*2} L 1 0"))
Plotting the data -
#collapse
base = alt.Chart(states).mark_geoshape(fill='#ededed', stroke='white').encode(text='name:N', longitude='lon:Q', latitude='lat:Q').project('albersUsa')
text = alt.Chart(states).mark_text().encode(text='name:N', longitude='lon:Q', latitude='lat:Q', tooltip=['name:N']).project('albersUsa')
spikes = alt.Chart(plot_data).mark_point(
fillOpacity=0.5,
fill='red',
strokeOpacity=1,
strokeWidth=1,
stroke='red',
).encode(
shape=alt.Shape('height', scale=None),
longitude='lon:Q',
latitude='lat:Q',
).project('albersUsa').properties(width=1200, height=900)
(base+text+spikes).configure_view(stroke=None)
Same data processing steps as above but we don't restrict ourselves to one date. That's the idea of the interactive plot. As we move the thumb we change the date for which the data is displayed.
covid = raw_data.copy()
# Making this on 27th October so limiting data till 27th becasue publishing it on website will take more time and experiments with the looks and interactivity
covid = covid[covid['date']<='2020-10-27']
# Albers USA projection does not support the following.
covid = covid[~((covid['state'] == "Puerto Rico")|(covid['state'] == "Virgin Islands")|(covid['state'] == "Guam")|(covid['state'] == "Northern Mariana Islands"))]
covid.loc[covid['county'] == 'New York City','fips'] = 1
# Getting per day statistics on CASES
covid = covid.assign(daily=covid.groupby('fips')['cases'].diff())
#covid = covid[covid['date'] > '2020-05-01'] # Working with data from May
# Getting past 2 weeks data for cases per given date - sum of cases in past 2 weeks
covid = covid.assign(past_2_weeks = covid.groupby('fips')['daily'].apply(lambda case: case.rolling(window=14).sum().shift(1)))
# Merging population estimate data - we could have done it earlier but what the heck!
## Getting census data
census = pd.read_csv('co-est2019-alldata.csv')
census = census[['STATE', 'COUNTY', 'STNAME', 'CTYNAME','POPESTIMATE2019']] # Keeping only 2019 population estimates
## Constructing FIPS from STATE and COUNTY codes in census itself
census['STATE'] = census.STATE.astype(str)
census['COUNTY'] = census.COUNTY.astype(str)
census['STATE'] = census.STATE.str.pad(2, fillchar='0')
census['COUNTY'] = census.COUNTY.str.pad(3, fillchar='0')
census = census.assign(fips = census.STATE + census.COUNTY)
census.fips = census.fips.astype(int)
nyc = census[(census['STATE'] == '36') & (census['COUNTY'].isin(['005', '047', '061', '081', '085']))]
nyc_census = pd.DataFrame([[36, 1, 'New York', 'New York City',nyc['POPESTIMATE2019'].sum(),1]], columns=['STATE','COUNTY','STNAME','CTYNAME','POPESTIMATE2019','fips'])
census = pd.concat([census, nyc_census])
census = census.drop(columns=['STATE','COUNTY','STNAME','CTYNAME'])
## Merging the census data with covid data on FIPS
covid = covid.merge(census, on='fips', how='inner')
## Getting the per capita data
covid = covid.assign(past_2_weeks_per_capita = covid.past_2_weeks/covid.POPESTIMATE2019*1000)
covid = covid[covid['past_2_weeks_per_capita'].notna()]
# Getting latitude and longitude
covid = covid.merge(counties, on='fips', how='inner')
covid = covid.drop(columns=['county', 'geometry', 'state', 'fips', 'cases', 'deaths', 'daily', 'past_2_weeks', 'POPESTIMATE2019', 'name'])
# Selecting dates when past two weeks sum of cases peaked
#bonus = covid[covid['past_2_weeks_per_capita']>2.5]
print(covid.info())
covid.head()
To play with the interactive chart online in this page, I have restricted the data by filtering data where the per capita last 2 weeks cases were more than 2.5
If you run it locally then you can run the whole thing using the data_server.
covid = covid.assign(height = covid['past_2_weeks_per_capita'].apply(lambda x: f"M -1 0 L 0 -{x*2} L 1 0"))
covid = covid[covid['past_2_weeks_per_capita']>2.5]
print(covid.info())
To be able to view the interactive chart, I have uploaded the data that Altair needs as a json file and I will directly pass the url to the Chart class.
url = "https://raw.githubusercontent.com/armsp/covidviz/master/assets/virus_surge_data.json"
#collapse
def timestamp(t):
return pd.to_datetime(t).timestamp() * 1000
slider = alt.binding_range(name='till_date:', step=1 * 24 * 60 * 60 * 1000, min=timestamp(min(bonus['date'])), max=timestamp(max(bonus['date'])))
day = alt.selection_single(bind=slider, name="slider", fields=['date'], init={'date':timestamp(min(bonus['date']))})
spikes = alt.Chart(url,).mark_point(
fillOpacity=0.3,
fill='red',
strokeOpacity=1,
strokeWidth=1,
stroke='red'
).encode(
latitude="lat:Q",
longitude="lon:Q",
shape=alt.Shape("height:N", scale=None),
# size='cases:Q'
).project(
type='albersUsa'
).add_selection(day).transform_filter(
"(year(datum.date) == year(slider.date[0])) && (month(datum.date) == month(slider.date[0])) && (date(datum.date) == date(slider.date[0]))"
)
(base+text+spikes).properties(width=1500, height=800, padding={'top': 225, 'bottom': 5}).configure_view(stroke=None)